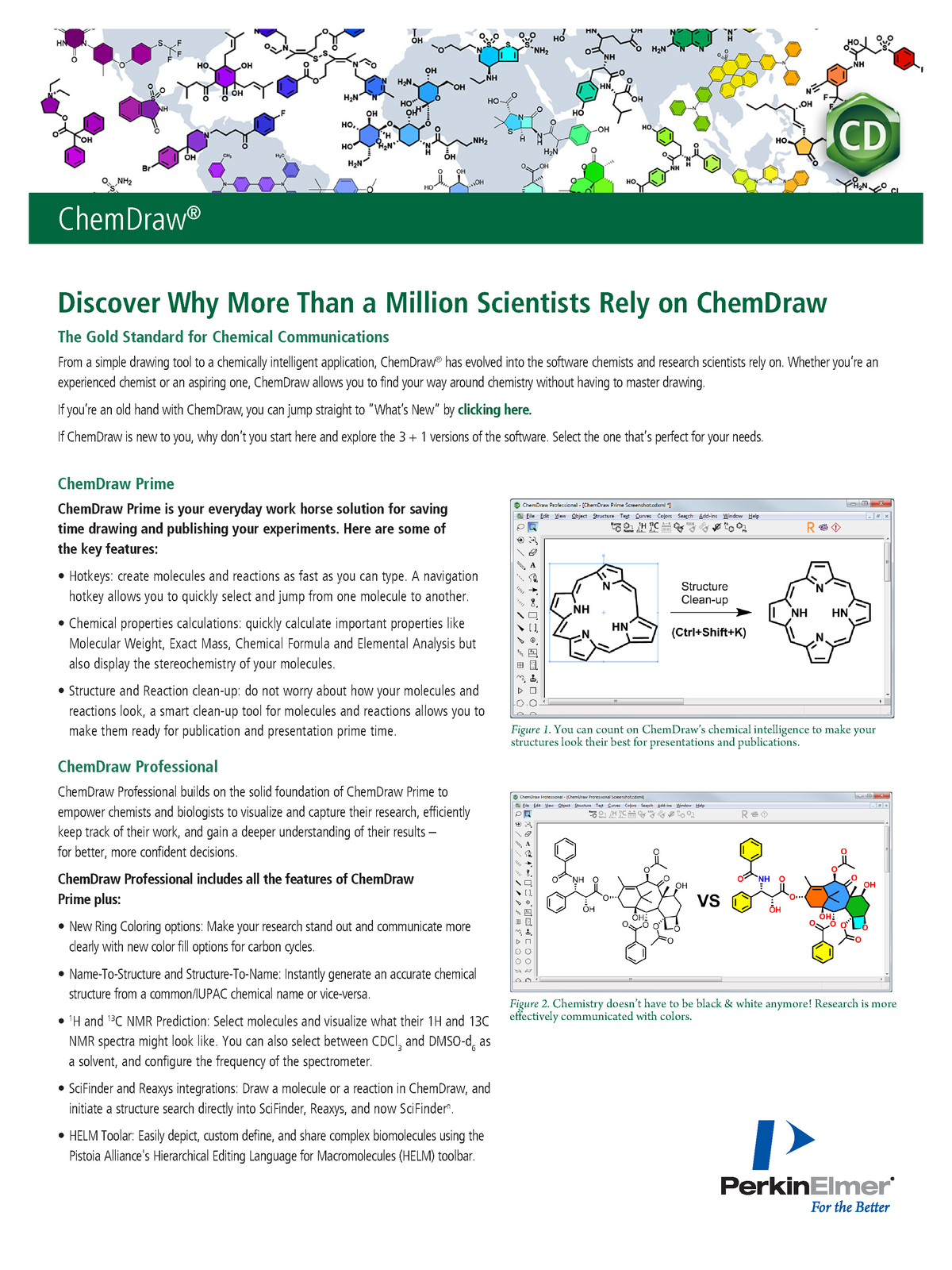

Try that for your version - if no luck I'll check on one of my other processing stations tomorrow which has version 9 installed. Then from the text toolbar (by default is usually at the bottom of the screen, but if not shown go to >View>Toolbars>Text) change the font size to something small enough to fit on the screen. 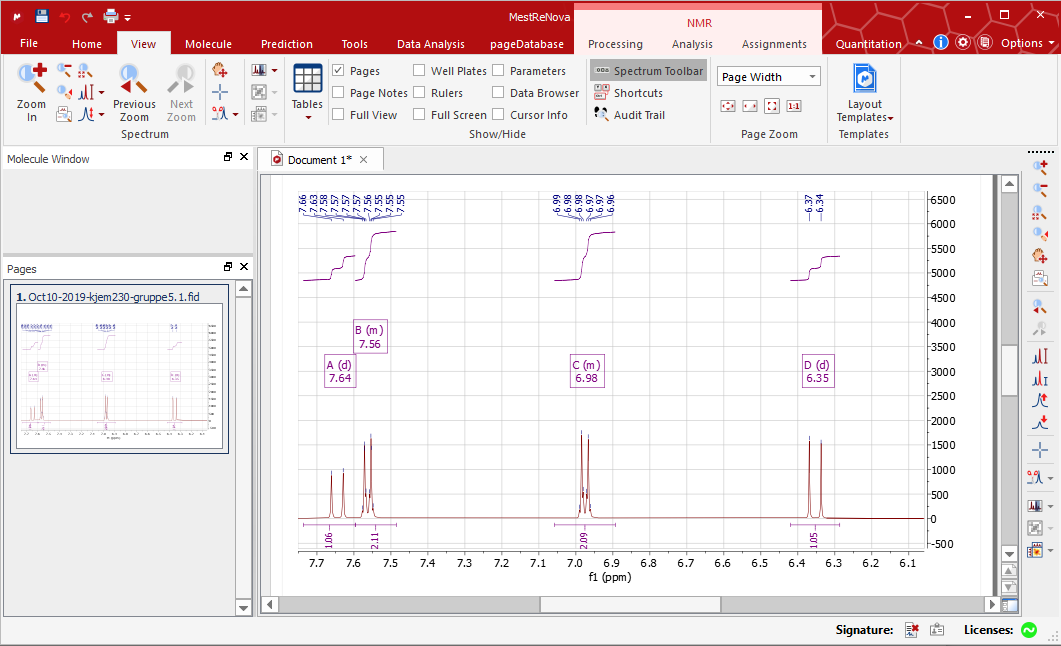 
Make sure under the geometry tab, the Auto box is selected for size. Left click the parameters (which should have the green selected boxes shown) and choose properties. ADVANCED Accurate prediction of 1 H and 13 C NMR spectra from a chemical structure and others ADVANCED Automatic confirmation of structure identity based on NMR and/or LC/GC/MS data. BASIC Carry out the analysis of various optical spectroscopy data. 2 Examples would include Brukers CONVDTA command and mNovas. BASIC Process, analyze and report your Mass Spec and/or Chromatographic data. 
You need to resize the text to a smaller font size directly from SpinWorks in PDF format (requires Adobe Acrobat reader) or in Word format. Left click and hold to define a box within which you will have your peaks pickedģ)how on earth do you get the parameter box to fit onto the page? Ive managed to get it up but it is about three times larger than my spectra and rescaling using the little green buttons leaves it the same size but just cuts loads out. >Analysis >Peak Picking >Manual Threshold (or just type K) >View >Zoom >Manual Zoom (or just type M, or click the manual zoom button)Ģ)Set a threshold so that only peaks above a certain intensity will automatically label. Testing this on version 9 will have to wait until tomorrow, but on version 8 (which I have in front of me, but which has limited differences to version 9 by memory)ġ)Rescale the chemical shift axis to focus only on the relevant sections of my spectra.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
